Demo of a KDE plot beside timeseries set (Interactive)¶
%pylab inline
#%matplotlib inline
#import matplotlib.pyplot as plt
import mpld3
from mpld3 import plugins
#mpld3.enable_notebook()
#%matplotlib notebook
#
import pysd
import numpy as np
import pandas as pd
import seaborn
Populating the interactive namespace from numpy and matplotlib
Load the model using PySD¶
The model is a basic, 1-stock carbon bathtub model
model = pysd.read_vensim('../../models/Climate/Atmospheric_Bathtub.mdl')
print(model.doc)
Real Name Py Name Subscripts Units 0 Emissions emissions None None
1 Excess Atmospheric Carbon excess_atmospheric_carbon None None
2 FINAL TIME final_time None Month
3 INITIAL TIME initial_time None Month
4 Natural Removal natural_removal None None
5 Removal Constant removal_constant None None
6 SAVEPER saveper None Month
7 TIME STEP time_step None Month
8 Time time None None
Limits Type Subtype Comment
0 (nan, nan) Constant Normal None
1 (nan, nan) Stateful Integ None
2 (nan, nan) Constant Normal The final time for the simulation.
3 (nan, nan) Constant Normal The initial time for the simulation.
4 (nan, nan) Auxiliary Normal None
5 (nan, nan) Constant Normal None
6 (0.0, nan) Auxiliary Normal The frequency with which output is stored.
7 (0.0, nan) Constant Normal The time step for the simulation.
8 (nan, nan) None None Current time of the model.
Generate a set of parameters to use as input¶
Here, drawing 100 constant values for the Emissions parameter from
an exponential distribution
n_runs = 100
runs = pd.DataFrame({'Emissions': np.random.exponential(scale=10000, size=n_runs)})
runs.head()
| Emissions | |
|---|---|
| 0 | 13139.288132 |
| 1 | 10414.796999 |
| 2 | 8505.605002 |
| 3 | 55251.743520 |
| 4 | 3650.497881 |
Run the model with the various parameters¶
result = runs.apply(lambda p: model.run(params=dict(p))['Excess Atmospheric Carbon'],
axis=1).T
result.head()
| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | ... | 90 | 91 | 92 | 93 | 94 | 95 | 96 | 97 | 98 | 99 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | ... | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
| 1 | 13139.288132 | 10414.796999 | 8505.605002 | 55251.743520 | 3650.497881 | 7490.206883 | 5908.429778 | 4526.126740 | 3969.536964 | 42777.645014 | ... | 9654.733184 | 16864.050543 | 15725.059944 | 7426.330819 | 10563.284279 | 5586.831254 | 24433.431913 | 13290.730858 | 6245.476841 | 11057.440643 |
| 2 | 26147.183383 | 20725.446028 | 16926.153954 | 109950.969604 | 7264.490783 | 14905.511697 | 11757.775259 | 9006.992213 | 7899.378559 | 85127.513578 | ... | 19212.919037 | 33559.460581 | 31292.869289 | 14778.398329 | 21020.935715 | 11117.794196 | 48622.529506 | 26448.554408 | 12428.498913 | 22004.306879 |
| 3 | 39024.999681 | 30932.988567 | 25262.497417 | 164103.203428 | 10842.343756 | 22246.663463 | 17548.627285 | 13443.049031 | 11789.921738 | 127053.883456 | ... | 28675.523031 | 50087.916518 | 46705.000540 | 22056.945165 | 31374.010637 | 16593.447508 | 72569.736124 | 39474.799721 | 18549.690765 | 32841.704452 |
| 4 | 51774.037816 | 41038.455680 | 33515.477445 | 217713.914913 | 14384.418199 | 29514.403711 | 23281.570790 | 17834.745281 | 15641.559485 | 168560.989636 | ... | 38043.500985 | 66451.087897 | 61963.010478 | 29262.706532 | 41623.554810 | 22014.344287 | 96277.470675 | 52370.782582 | 24609.670698 | 43570.728050 |
5 rows × 100 columns
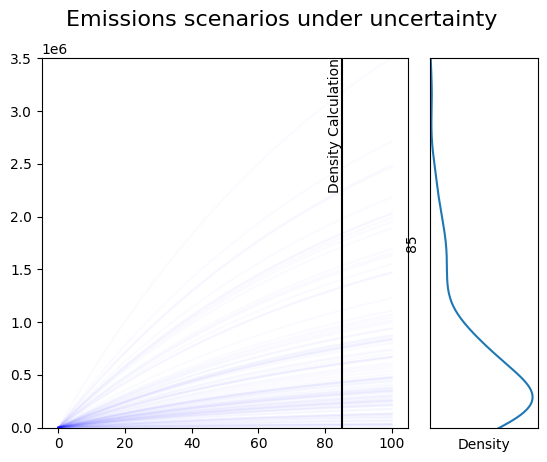
Draw a static plot showing the results, and a marginal density plot¶
This would be for making graphics for a publication, where you don’t want an interactive view, but you want fine control over what the figure looks like.
This is relatively simple, because we rely on the plotting library
seaborn to generate the KDE plot, instead of working out the
densities ourselves.
import matplotlib.pylab as plt
# define when to show the density
density_time = 85
# left side: plot all traces, slightly transparent
plt.subplot2grid((1,4), loc=(0,0), colspan=3)
[plt.plot(result.index, result[i], 'b', alpha=.02) for i in result.columns]
ymax = result.max().max()
plt.ylim(0, ymax)
# left side: add marker of density location
plt.vlines(density_time, 0, ymax, 'k')
plt.text(density_time, ymax, 'Density Calculation', ha='right', va='top', rotation=90)
# right side: gaussian KDE on selected timestamp
plt.subplot2grid((1,4), loc=(0,3))
seaborn.kdeplot(y=result.loc[density_time])
plt.ylim(0, ymax)
plt.yticks([])
plt.xticks([])
plt.xlabel('Density')
plt.suptitle('Emissions scenarios under uncertainty', fontsize=16);
Interactive plot using python backend¶
The following would be for lightweight exploration
import matplotlib as mpl
from ipywidgets import interact, IntSlider
sim_time = 200
slider_time = IntSlider(description = 'Select Time for plotting Density',
min=0, max=result.index[-1], value=1)
@interact(density_time=slider_time)
def update(density_time):
ax1 = plt.subplot2grid((1,4), loc=(0,0), colspan=3)
[ax1.plot(result.index, result[i], 'b', alpha=.02) for i in result.columns]
ymax = result.max().max()
ax1.set_ylim(0, ymax)
# left side: add marker of density location
ax1.vlines(density_time, 0, ymax, 'k')
ax1.text(density_time, ymax, 'Density Calculation', ha='right', va='top', rotation=90)
# right side: gaussian KDE on selected timestamp
ax2 = plt.subplot2grid((1,4), loc=(0,3))
seaborn.kdeplot(y=result.loc[density_time], ax=ax2)
ax2.set_ylim(0, ymax)
ax2.set_yticks([])
ax2.set_xticks([])
ax2.set_xlabel('Density')
plt.suptitle('Emissions scenarios under uncertainty', fontsize=16);
interactive(children=(IntSlider(value=1, description='Select Time for plotting Density'), Output()), _dom_clas…
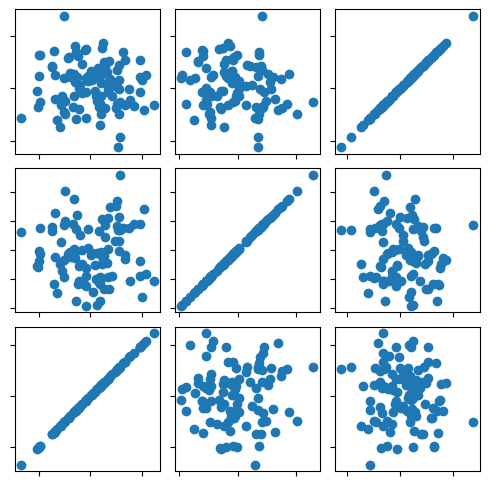
Interactive figure with javascript background¶
This script would prepare interactive graphics to share on a webpage, independent of the python backend.
fig, ax = plt.subplots(3, 3, figsize=(6, 6))
fig.subplots_adjust(hspace=0.1, wspace=0.1)
ax = ax[::-1]
X = np.random.normal(size=(3, 100))
for i in range(3):
for j in range(3):
ax[i, j].xaxis.set_major_formatter(plt.NullFormatter())
ax[i, j].yaxis.set_major_formatter(plt.NullFormatter())
points = ax[i, j].scatter(X[j], X[i])
plugins.connect(fig, plugins.LinkedBrush(points))